Director
University of Maryland, Maryland Technology Enterprise Institute (Mtech)

William E. Bentley is the Robert E. Fischell Distinguished Chair of Engineering, the Inaugural Director of the Robert E. Fischell Institute for Biomedical Devices, and the Director of the Maryland Technology Enterprise Institute. He is also appointed to the Department of Chemical and Biomolecular Engineering at the University of Maryland, College Park and the Institute for Bioscience and Biotechnology Research. At Maryland since 1989, Dr. Bentley has focused his research on the development of molecular tools that facilitate the expression of biologically active proteins, having authored over 400 related archival publications. His recent interests interface biology with electronics, establishing electrogenetics as a means for electronic control of biological systems.
He is a fellow of AAAS, ACS, AIMBE, and the American Academy of Microbiology. He has served on advisory committees for the NIH, NSF, DOD, DOE, FDA, USDA, and several state agencies and has mentored over 50 PhDs and 25 postdocs, many now in leadership roles within industry (24), federal agencies (5) and academia (26). He co-founded a protein manufacturing company, Chesapeake PERL, based on insect larvae as mini bioreactors.
University of Maryland, Maryland Technology Enterprise Institute (Mtech)
University of Maryland, Robert E. Fischell Institute for Biomedical Devices
University of Maryland, Fischell Department of Bioengineering
University of Maryland, Glenn L. Martin Institute of Technology
University of Maryland, Glenn L. Martin Institute of Technology
University of Maryland, Department of Chemical and Biomolecular Engineering
Ph.D. in Chemical Engineering
University of Colorado, Boulder
Master of Engineering in Chemical Engineering
Cornell University
Bachelor of Science in Chemical Engineering
Cornell University

ECI Conference on Biochemical and Molecular Engineering XXIII: Accelerating Biotech Solutions to aid a Changing World, Sponsored by Amgen, Inc., in memory of James E. Bailey

For supporting the growth and development of students, post-docs, and early career faculty; motivating and empowering mentees to pursue and achieve their goals, and helping mentees build a network of professional relationships to strengthen their careers.

This annual award recognizes individuals for their contributions to the field and to the practice of biochemical engineering through their position in industry or academia as exemplified by Professor Wang in his 50 years of contributions.

Distinguished University Professors will have been recognized nationally and internationally for the importance of their scholarly and/or creative achievements. They will also have demonstrated the breadth of interest characteristically encompassed by the traditional role of scholar, teacher, and public servant. In addition, they will have brought distinction to the University of Maryland as a result of those activities.

The Marvin J. Johnson Award in Microbial and Biochemical Technology, to recognize outstanding research contributions to microbial and biochemical technology. The Award is presented by the Division of Biochemical Technology at the Spring meeting of the American Chemical Society. The Award was established in 1978 and is currently sponsored by Pfizer.

This award is given to recognize individuals who have made one or more outstanding research contributions in industrial microbiology and/or biotechnology. These contributions should be of exceptional merit, reflecting an independence of thought and originality that adds appreciably to scientific knowledge. Activities such as journal editing, organizing and chairing conferences, and serving scientific societies in official capacities also may be considered when judging research contributions. However, the most important factor in selecting nominees for this Award is research accomplishments.

The American Chemical Society (ACS) Fellows Program was created by the ACS Board of Directors in December 2008 to recognize members of ACS for outstanding achievements in and contributions to science, the profession, and the Society.

Recognizes an individual's outstanding chemical engineering contribution in the food, pharmaceutical and/or bioengineering field, of fundamental nature and/or of practical significance to industry and industrial practice. These contributions may have been made in industry, government, or academic settings, or with other organizations.

The American Institute for Medical and Biological Engineering (AIMBE) is a non-profit organization headquartered in Washington, D.C., representing 50,000 individuals and the top 2% of medical and biological engineers. In addition, AIMBE represents academic institutions, private industry, and professional engineering societies. AIMBE was founded in 1991 and its current vision is to provide leadership and advocacy in medical and biological engineering for the benefit of society.

Over the last 50 years, 2,700 distinguished scientists have been elected to the Academy. Fellows are elected through a highly selective, annual, peer review process, based on their records of scientific achievement and original contributions that have advanced microbiology. A Committee on Elections, consisting of Fellows of the Academy who are elected by the membership, reviews all nominations for Fellowship and recommends to the Board of Governors what action should be taken.

A member whose efforts on behalf of the advancement of science or its applications are scientifically or socially distinguished and who has been a continuous member for the four year period leading up to the year of nomination, may, by virtue of such meritorious contribution be elected a Fellow by the Council. Examples of areas in which nominees may have made significant contributions are research; teaching; technology; services to professional societies; administration in academe, industry, and government; and communicating and interpreting science to the public. Fellows are elected annually by the AAAS Council from the list of approved nominations from the Section Steering Committees.

AIMBE's College of Fellows includes around 1,500 individuals who have made significant contributions to the medical and biological engineering (MBE) community whether in academia, industry, or government and their contributions to MBE research, industry practice, and education have transformed the world.
Our lab is developing methodologies to interrogate and control molecular signaling, both inside and outside of cells. We use the tools of metabolic engineering and synthetic biology to rewire genetic circuits that enable these actions. We are particularly interested in connecting biology to electronics to facilitate “programming” of biological function. We have discovered that redox provides an almost unique capability for this information transfer as it is a fundamental communication modality within biology and is directly accessed via electronics. Our lab mainly focuses on bacterial systems and their interactions with eukaryotic cells, we exploit the molecular components of bacterial quorum sensing, a cell-cell signaling phenomenon our group originally helped to discover.

From Left to Right: Chen-Yu Chen, Varun Kochar, Joshua Dayie, Alex Compean, Divya Muthusamy, David Boegnar, Futoon Al-Jirafi, Rahma Zakaria, Chen-Yu Tsao, Briahnah Streeter, Noah Cauchi, Monica Chu, Ben Wu, William Bentley

From Left to Right: William Bentley, Chen-Yu Chen, Eric VanArsdale, Sally Wang, Chen-Yu Tsao, Jinyang Li, Dana Motabar, Kayla Chun, Monica Chu, Futon Al-Jirafi, Rahma Zakaria (not pictured, Nick Pirolli)

From Left to Right: Chen-Yu Chen, Eric VanArsdale, Dana Motabar, William Bentley, Sally Wang, So Hyun Ahn, Kayla Chun, Monica Chu, Rahma Zakaria (not pictured, Kristina Stephens, Chen-Yu Tsao, Ashley Chapin, Julie Pitzer)

From Left to Right: Kristina Stephens, So Hyun Ahn, Dana Motabar, Kayla Chun, Ashley Chapin, Jinyang Li, Chen-Yu Tsao, Eric VanArsdale, Sally Wang, Rahma Zakaria, Chen-Yu Chen, Julie Pitzer

From Left to Right: Tanya (Gordonov) Tschirhart, Wu Shang, Ryan McKay, William Bentley, Amin Zargar, Naren Bhokisham, David Quan, Melissa Rhoads, Jess Terrell, Kristina Stephens, Hana Ueda, Chelsea Virgile, Michelle Brookstaver

From Left to Right: David Quan, Haig Pakhchanian, Amin Zargar, Naren Bhokisham, Tanya Gordonov, Niketa Jani, Milad Emamian, Jessica Terrell, William Bentley, Chen-Yu Tsao, Hsuan-Chen Wu, Nathan Barber, Hana Ueda, Melissa Rhoads, Chelsea Virgile, Steve Graff, John Kerwin, Kevin Knapstein, Darryll Sampey


A selective list of publications that Dr. Bentley has authored/co-authored in the past few years. For a complete list of Dr. Bentley's publications, please visit his Google Scholar profile.
Latest news and updates from the Bentley Research Group and the Fischell Institute for Biomedical Devices at the University of Maryland.

Dr. Reza Ghodssi, University of Maryland Distinguished University Professor, among keynote speakers at AUKUS Forum
Read more
How a high-impact strategy launched by a transformative, $220 million investment in UMD is creating new, tangible momentum for engineering progress on important societal challenges.
Read more
The MPower Fellowship supports recent graduates from the University of Maryland's College Park and Baltimore campuses in transforming biomedical or medical device innovations into market-ready prod...
Read more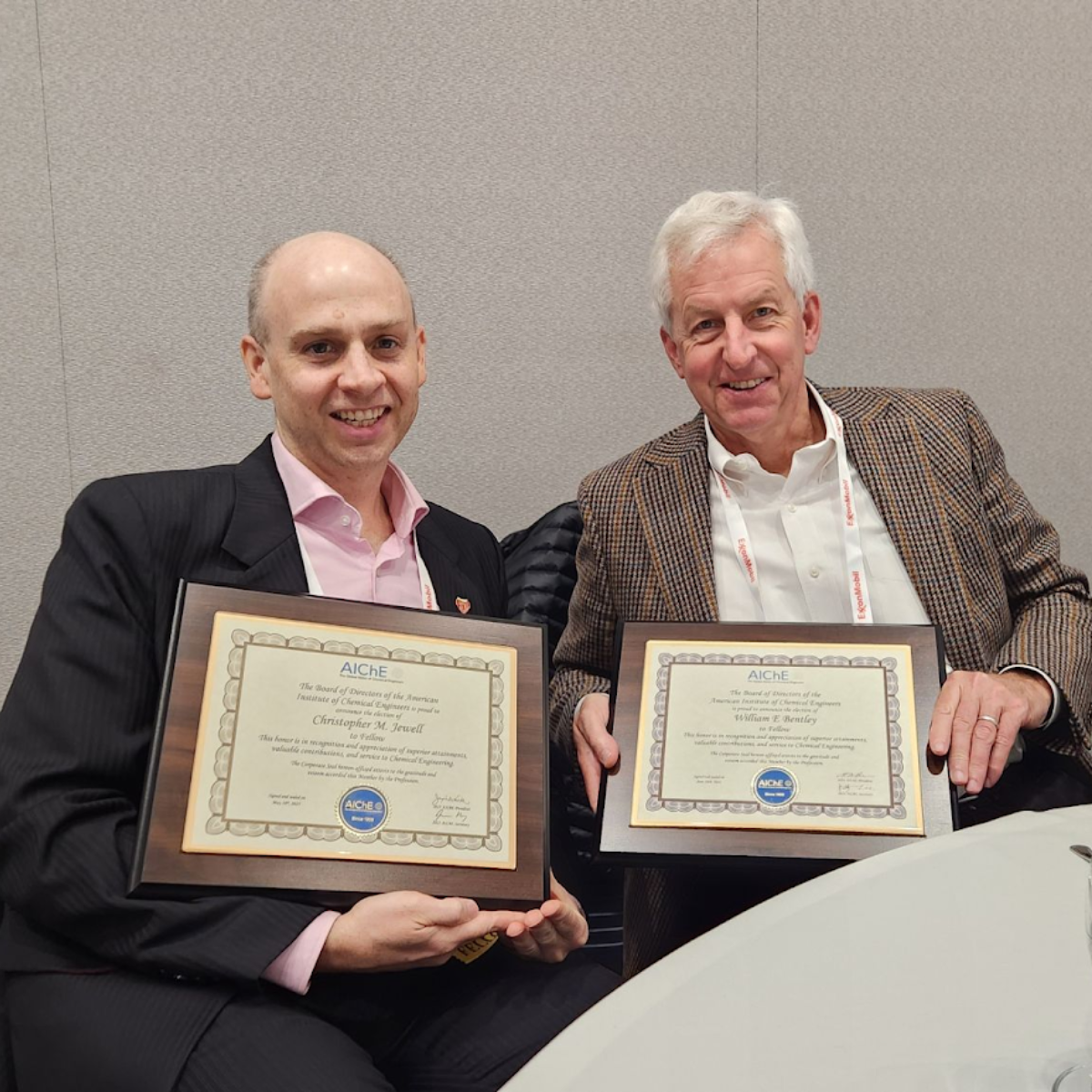
Bill Bentley and Chris Jewell were recognized as AIChE Fellows at the 2025 AIChE Annual Meeting in Boston.
Read more
Chemical engineers design solution to fast-charge lithium-ion batteries for EVs.
Read more
At BMES 2025, UMD Ph.D. student Lauren A. Gomes presented 3D neural tissue research aimed at modeling neuroplasticity and guiding future therapies.
Read more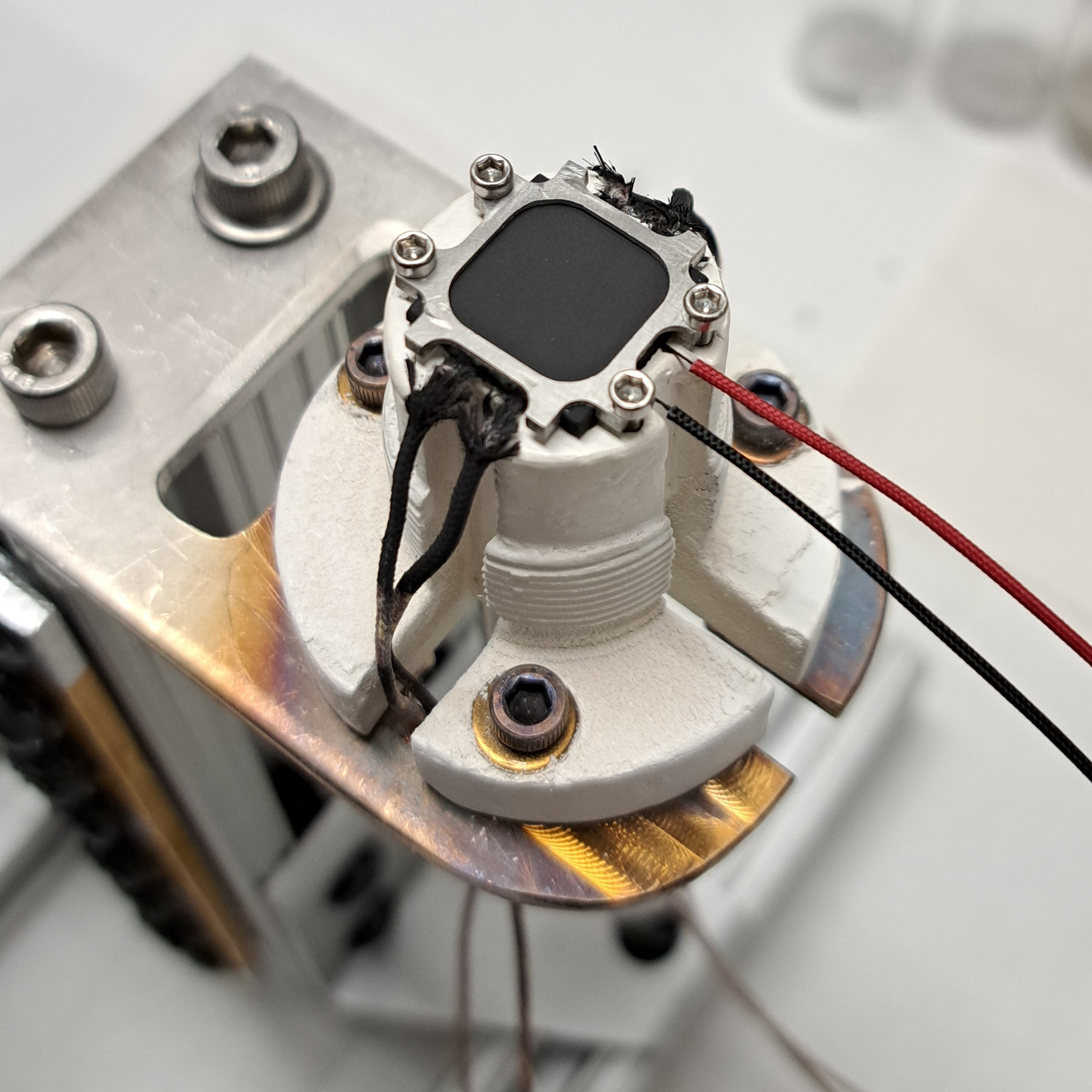
UMD researchers develop improved sensors for measuring heat flux in real-time with more precision in extreme environments.
Read more
The Jewell Research Lab, was recently published in three top journals: Proceedings of the National Academy of Sciences (PNAS), Cell Press Molecular Therapy, and Biomaterials.
Read more
Engineering PhD student Sydney Overton advancing research while supporting peers
Read more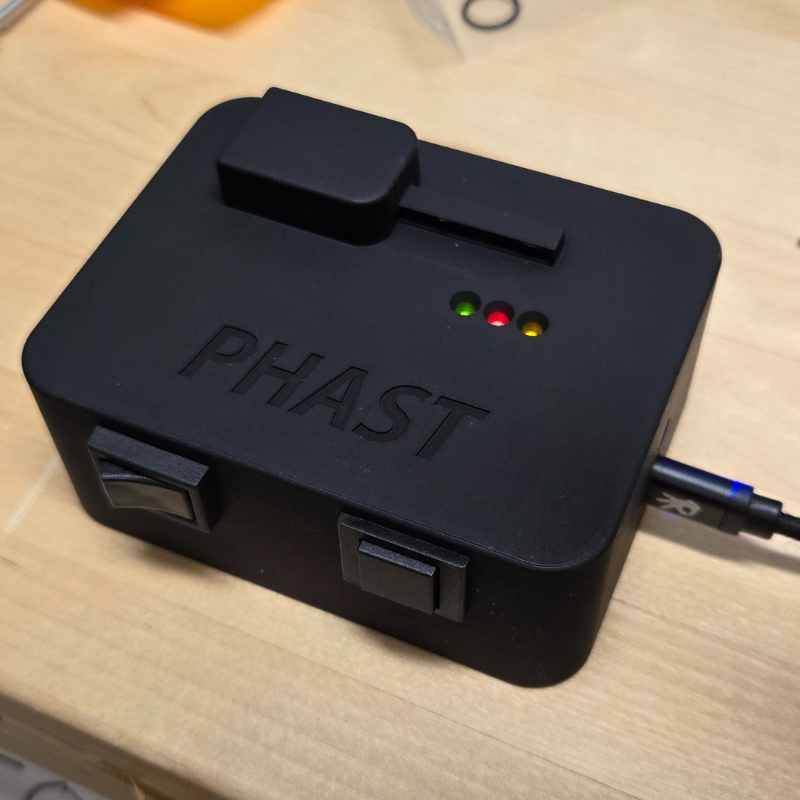
PHAST is a portable device that could help doctors quickly assess a patient's risk of multiple organ failure after severe trauma.
Read more
UMD mechanical engineering faculty member is recognized for his contributions to additive manufacturing.
Read more
Ashly Cifuentes is the Fischell Institute Student Assistant where she supports staff, events, ensuring a safe research environment, and keeping track of packages.
Read more
Annual Convocation event celebrates academic and service excellence and achievement.
Read more
UMD Researchers Featured Front Cover on Biomaterials Science for Advances in Drug Delivery
Read more
The Research Symposium is where Fischell Institute interns present the projects they worked on and reflected on their experience in the program.
Read more
Two BIOE Ph.D. students were named Fischell Fellows at the eighth annual EPIC Retreat.
Read more
Appointment effective September 1, 2025
Read more
Hannah Zierden will lead an investigation to understand biological processes occurring in mucosal barriers, a channel for targeted local drug delivery.
Read more
Projects promote multidisciplinary collaboration and innovative research
Read more
Honor recognizes excellence, impact, and significant contribution
Read moreWe emphasize quantitative understanding of biology and biological systems at the cellular and molecular levels. If we can understand biology’s design concepts and if we can recapitulate them mathematically, we could better understand the progression of disease and develop better medicines. We may even be in a position to create “smart” devices that interrogate biology and take action when needed.
This course reinforces traditional models of macromolecular function and cell biology with the quantitative approach typical in engineering in order to build a more profound, intuitive understanding of cell biology and its application for improving human health. We address design and analysis of biomolecular structure and function, molecular interactions, enzymes, metabolic pathways, signal transduction, synthetic genetic circuits, and cell systems and their cultivation.
Presentation of techniques for characterizing and manipulating non-linear biochemical reaction networks. Advanced topics to include mathematical modeling of the dynamics of biological systems; separation techniques for heat sensitive biologically active materials; and rate processes in cellular and biomolecular systems. Methods are applied to current biotechnological systems, some include: recombinant bacteria; plant, insect and mammalian cells; and transformed cell lines.
A combination of lectures and discussions covering biology from a utilization perspective, and lectures on illustrative mathematical models that capture the essences of characteristics of living entities. The biology material will focus on: distinguishing engineering from biological science, principles form the sciences applicable to biology, typical biological responses to environmental stimuli, scaling of biological responses, and different means to utilize living entities.
Dr. Bentley strives to make himself as available as possible to students and faculty alike. If you'd like to meet with him or just get in touch, please use the contact information to the right. Email is the most practical form of communication and will typically yield the fastest response.
Dr. Bentley does not have posted office hours. Please email him to schedule a time to meet.
Dr. Bentley's office is located on the fifth floor of A. James Clark Hall.
5102 A. James Clark Hall
University of Maryland, College Park, MD 20742
Dr. Bentley has two labs on campus, also located in Clark Hall.
A. James Clark Hall
University of Maryland, College Park, MD 20742